Engineering Protein Localization
Most functions of a cell are carried out by proteins. Our ability to manipulate proteins and their functions has opened up numerous possibilities for treating diseases at the molecular level. For decades, drug development has largely focused on blocking or activating the function of disease-related proteins to correct the pathology. A classic example for protein inhibition is the use of small-molecule inhibitors that bind to the protein and block its function, or even more effective in some cases, Targeted Protein Degradation (TPD) technologies that eliminate disease-causing proteins by tagging them for destruction. But what if the problem is not with the protein's activity, but rather where it is located? Targeted Protein relocalization technologies focus on changing the localization of a particular protein, to establish protein function where it is most needed.
Subcellular Mislocalization of Proteins in Human Diseases
Cells are organized into distinct membrane-bound compartments known as organelles, where various metabolic activities and reactions occur. These include the nucleus, endoplasmic reticulum (ER), Golgi apparatus, mitochondria, lysosomes, and peroxisomes. Think of a cell as a small town with different buildings where different activities take place. In each building we find people taking care of different responsibilities. Proteins, like people with specific jobs, are often precisely localized to carry out their functions effectively, just as an experimental scientist must be precisely located in the lab to perform their work.
Protein mislocalization is a frequent cause of disease, emphasizing how crucial cellular location is for proper function. For example:
Cancer: Mislocalized tumor suppressors, such as p53 that should reside in the nucleus to regulate gene expression and prevent uncontrolled cell growth, are major cancer contributors.
Neurodegenerative Diseases: Accumulation of mislocalized TDP43 and FUS proteins in the cytosol (instead of the nucleus) are key contributors to conditions like Amyotrophic Lateral Sclerosis (ALS) and Frontotemporal Lobar Degeneration (FTLD).
Retinitis Pigmentosa: The rhodopsin protein, essential for vision, is retained in the ER instead of reaching the plasma membrane where it functions, which leads to retinal degeneration.
Cystic Fibrosis: Mutant Cystic Fibrosis Transmembrane Conductance Regulator (CFTR) proteins accumulate in the ER and do not reach the cell surface, where they regulate ion transport across membranes.
Inhibiting or activating protein function approaches fall short when dealing with diseases caused by proteins being in the wrong place rather than simply malfunctioning. This is where targeted protein relocalization has major relevance.
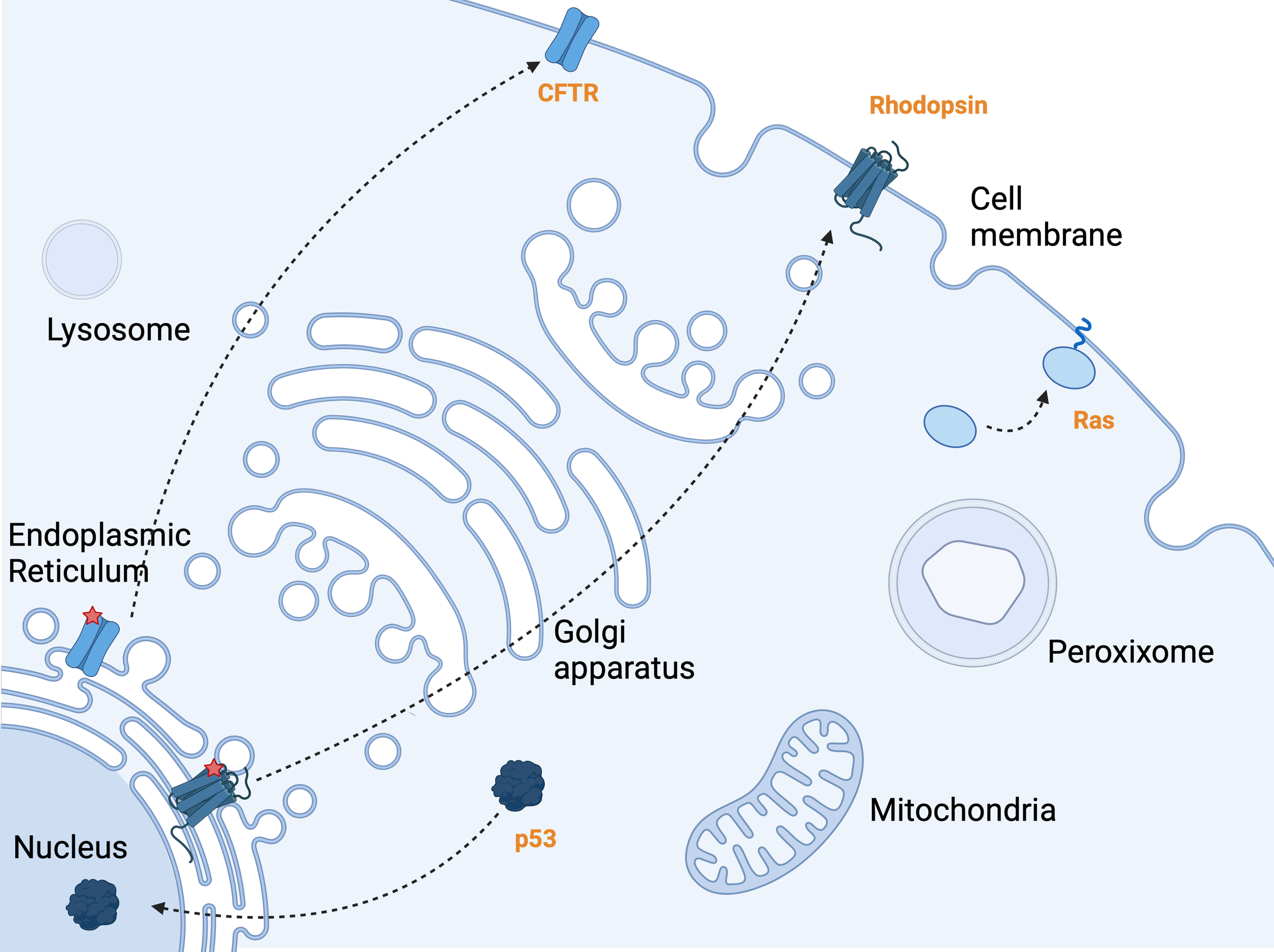
A cell with various compartments, highlighting subcellular protein trafficking.
p53 moves from the cytosol to the nucleus, where it functions as a tumor suppressor by regulating gene expression. Rhodopsin, a membrane protein, is trafficked to the cell surface, while a mutant version remains stuck in the ER. CFTR follows a similar pattern, with normal trafficking to the cell surface and mutant CFTR accumulating in the ER, representing cystic fibrosis pathology. Ras, an oncogene, relocates from the cytosol to the cell membrane, where it becomes active through lipid molecule addition, triggering signaling pathways associated with cancer progression.
Targeting Mechanisms for Protein Localization
The subcellular distribution of many disease-relevant proteins can be influenced by binding to other biomolecules that stabilize the protein in one compartment or that traffic the protein to its final destination. These include coat proteins that load proteins into transport vesicles or importins that import proteins into the nucleus.
Protein localization can also be influenced by a wide variety of protein modifications, called post-translational modifications, as these additions occur after the protein is built or translated. These include: phosphorylation, the addition of a phosphate group; acetylation, the addition of an acetyl group; ubiquitylation, the addition of a ubiquitin protein or lipidation, th addition of a lipid molecule.
When these mechanisms go wrong—either the interaction with transport proteins or the protein modifications—proteins end up in the wrong cellular compartment. Accordingly, manipulation of the upstream regulatory processes has the potential to restore the correct location and function of aberrantly localized proteins. From a chemical and pharmacological perspective, the action of many proteins involved in these transport pathways can be modulated by small molecules.
Therapeutic Strategies to Correct Protein Localization
Some molecules designed to manipulate protein localization work to restore a protein’s proper placement, while others intentionally mislocalize proteins to inhibit their function. Strategies aimed at correcting mislocalized proteins operate at different levels, addressing protein folding, trafficking, and activity.
Correctors and Potentiators
One major cause of protein mislocalization is faulty folding due to genetic mutations. Proteins must fold into specific three-dimensional structures to function correctly, but mutations can disrupt this process, leading to misfolding and improper transport. Correctors are molecules that bind to these defective proteins, stabilizing their structure and promoting correct folding and trafficking.
A well-known example comes from cystic fibrosis treatments, where correctors such as lumacaftor, tezacaftor, and vanzacaftor help refold mutant CFTR proteins, allowing them to reach the cell surface. However, proper localization alone isn’t always enough—some misfolded proteins also suffer from impaired activity.
This is where potentiators come in. These molecules bind to the protein and enhance its function, making it work more efficiently. In cystic fibrosis, the potentiators ivacaftor and deutivacaftor boost the channel activity of CFTR proteins, helping restore ion transport in patients when combined with correctors.
Together, correctors and potentiators represent a powerful approach to addressing diseases caused by protein misfolding and trafficking defects. However, some of these molecules are only effective for certain protein mutations, so other cutting-edge strategies are emerging to manipulate protein localization even more efficiently.
Guiding Proteins with Light and Self-Localizing Small Molecules
Two innovative approaches, optogenetics and self-localizing ligands (SLLs), offer dynamic ways to control protein localization using light or small-molecule signals.
Optogenetics enables the control of protein function with light. Proteins are composed of sequences of amino acids, and those that localize in a particular compartment of the cell possess localization signals—specific amino acid sequences that dictate their transport. To draw a parallel, these signals function like a movie theater ticket, which, when read by the usher, determines where you are seated. Similarly, cellular trafficking proteins interpret localization signals to direct proteins to the correct location.
One such sequence, the Nuclear Localization Signal (NLS), directs proteins to the nucleus. Researchers have engineered a light-activated NLS, called OptoNLS, which remains hidden in the dark and is exposed upon light activation, allowing precise spatiotemporal control of protein localization.
Self-Localizing Ligands (SLLs) offer a chemical-based alternative. These small synthetic molecules spontaneously localize to specific subcellular regions in mammalian cells. They posses built-in small-molecule localization signals — chemical structures with certain chemical properties or which interact with cellular components in a way that directs them to a particular compartment. These motifs provide passive or active targeting capabilities, making them useful in protein localization strategies.
Researchers have developed SLLs that spontaneously localize to specific membrane compartments in mammalian cells. These ligands bind target proteins and rapidly relocate them from the cytoplasm to designated sites, effectively serving as synthetic protein translocators.
By expanding the library of, these tools are unlocking new possibilities for precisely controlling protein positioning—setting the stage for the next breakthrough: chemical magnets.
Chemical Magnets
A new class of small molecules, known as chemical magnets, takes a different approach: they bind to two different proteins that wouldn't normally interact, bringing them together in a controlled manner. By tethering these two proteins together, these molecules can modulate protein levels, stability, activity, and even their location within the cell.
One such molecule, NICE-01, was recently developed to relocate proteins from the cytoplasm to the nucleus. It works by binding to BRD4, a nuclear-localized protein, and simultaneously capturing FKBP12F36V, a cytoplasmic protein. This dual-targeting design enables the nuclear transport of FKBP12F36V and any proteins fused to it, offering a potential strategy for directing therapeutic proteins to precise cellular locations.
Building on this concept, researchers in 2024 introduced Targeted Relocation-Activating Molecules (TRAMs), an approach that harnesses shuttle proteins—proteins that naturally travel between the nucleus and cytoplasm—to redirect disease-associated proteins. In cellular models of ALS, TRAMs successfully moved FUS, a mislocalized protein, from the cytoplasm back to the nucleus, reducing the formation of harmful stress granules linked to neurodegeneration. Another application of TRAMs demonstrated their ability to transport protective proteins into axons, slowing axonal degeneration in neurons. This method mimics protective mutations found in certain individuals, offering potential for treating neurodegenerative disorders by ensuring key proteins are in the right place at the right time.
By designing small molecules that precisely manipulate protein location, chemical magnets are opening up new therapeutic possibilities—not just for neurodegenerative diseases but also for cancer, where they could force oncogenic proteins into the nucleus, enhancing targeted therapies.
Nanobody-Mediated Relocalization
Beyond small molecules, nanobody-based strategies are emerging as another promising tool for protein relocalization. Nanobodies are tiny antibody fragments derived from camelids (camels, llamas, and alpacas) that can bind with high specificity to target proteins. Their small size and adaptability make them ideal for engineering protein localization.
By attaching localization signals to nanobodies, researchers can create molecules that recognize and shuttle mislocalized proteins to their correct destinations. For example, a nanobody targeting a protein trapped in the cytoplasm could be engineered with a nuclear localization signal, actively redirecting it to the nucleus.
The modular nature of nanobody-based systems allows for customized therapeutic strategies, where different nanobodies can be paired with different localization signals depending on the needs of a patient. This makes them an exciting frontier in personalized medicine, offering tailored solutions for diseases caused by protein mislocalization.
Limitations and Challenges of Therapeutic Strategies for Protein Relocalization
While protein relocalization strategies hold great promise, several key challenges must be addressed before they can be widely applied in clinical settings:
Ensuring Specificity
A major challenge is achieving high specificity in the relocalization process. The human body contains thousands of different proteins, and many share similar structures or domains. Relocalization technologies must be precise enough to target only the disease-relevant protein while leaving similar proteins untouched.
This is particularly challenging for small molecule approaches, which often have off-target effects. Even targeted approaches like nanobodies or TRAMs must be extensively validated to ensure they don't inadvertently redirect unintended proteins, which could disrupt normal cellular functions and potentially cause new health problems.
Achieving Consistent Outcomes Across Cell Types
The internal organization and protein trafficking machinery can vary significantly between different cell types. A relocalization strategy that works perfectly in liver cells, for instance, might be ineffective in neurons or muscle cells. This variation presents a significant challenge for developing treatments for diseases that affect multiple tissues.
Additionally, the severity of protein mislocalization can vary between patients with the same disease, making it difficult to develop one-size-fits-all relocalization strategies. This highlights the potential need for personalized approaches based on individual protein mislocalization patterns.
Delivery Challenges
Getting relocalization molecules or proteins into the right cells presents another major hurdle. For small molecules, issues like poor solubility, rapid metabolism, or inability to cross cellular membranes can limit effectiveness. For protein-based approaches such as nanobodies, delivery is even more challenging as proteins typically cannot easily enter cells.
Various delivery systems like lipid nanoparticles, viral vectors, or cell-penetrating peptides are being explored, but each comes with its own set of challenges related to safety, manufacturing scalability, and targeting specificity.
Immune Responses
Protein-based relocalization strategies using engineered components like nanobodies or shuttle proteins might trigger immune responses, especially with repeated dosing. The human immune system is designed to recognize and eliminate foreign proteins, which could lead to reduced efficacy over time or even adverse reactions.
Strategies to minimize immunogenicity, such as humanization of nanobodies or using delivery systems that shield proteins from immune detection, are being developed but add another layer of complexity to these approaches.
Temporal and Quantitative Control
For optimal therapeutic effect, relocalization strategies may need to achieve the right level of protein in the right place at the right time. Too little relocalization might be ineffective, while too much might disrupt other cellular processes.
Developing systems with tunable activity, perhaps responsive to specific cellular conditions or external regulators, represents an important frontier in this field.
The Road Ahead
Protein relocalization strategies represent a paradigm shift in how we think about treating disease. Rather than simply blocking harmful activities or replacing missing functions, these approaches aim to restore normal cellular organization—addressing the disease at its organizational level. The potential for more precise interventions with fewer side effects makes this an extremely promising area for continued research and development.
As we gain deeper understanding of the complex choreography of proteins within cells and develop more sophisticated tools to manipulate this dance, we may be able to correct the cellular disharmony underlying many currently untreatable conditions.
Would you like to stay updated on the latest breakthroughs in biomedical science?
Subscribe to my blog and join me in exploring the next frontier of medicine!